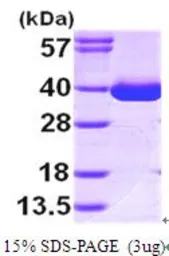
Human MDH1 protein, His tag (active)
Cat. No. GTX67090-pro
Cat. No. GTX67090-pro
-
ApplicationsFunctional Assay
-
SpeciesHuman