Knockdown Technology
|
 |
||
|
The challenges with RNA interference technology |
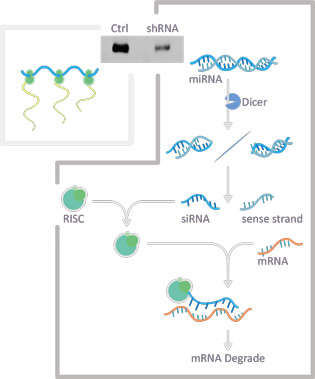
One challenge with siRNAs is that they are subject to degradation themselves, with only about 1% of the transfected duplex remaining two days after introduction. This means that large amounts must be transfected in order to maximize the chance of success. In addition, knockdown of the specific protein is transient. One way to address these two issues is to introduce into cells plasmids expressing short hairpin RNAs (shRNAs) that are converted into siRNAs by the endogenous RNAi machinery. This approach allows the persistent expression of siRNAs within the cell, thereby facilitating long-term knockdown of the target transcript. Various inducible or otherwise promoter-specific vectors can be used to create even more refined control of shRNA expression.
|
GeneTex engages RNA interference technology to validate antibody specificity |
At GeneTex, we use an shRNA-mediated RNAi protocol employing a lentiviral expression plasmid optimized for the expression of shRNAs. Once transported into the nucleus, the plasmid directs transcription of the shRNA that is subsequently processed like the miRNAs described above. This makes shRNA more efficient. In addition, the knockdown is more stable over time as the shRNA is constantly expressed. Interestingly, only a few copies of the shRNA-expressing vector need to be integrated into the cell's genome in order to maintain knockdown. This decreases the risk of off-target effects, which is a more prominent problem with transfected siRNAs at their higher introduced concentrations.
|
GeneTex complements these efforts with CRISPR/Cas9-mediated KO cell lines |
GeneTex will continue to expand its utilization of shRNA as a key component of its antibody validation program. In addition, we will incorporate other methodologies to guarantee the specificity of our products, including application of CRISPR/Cas9 technology. Made possible by our collaborative relationship with the genome editing technology company EdiGene, GeneTex will use lysates from EdiGene’s knockout (KO) cell lines to definitively demonstrate antibody specificity. At GeneTex, we understand the importance of quality reagents to your research. We are committed to providing best-in-class reagents and leading the research antibody industry in antibody validation.