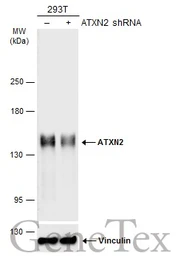
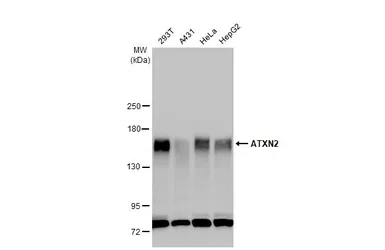
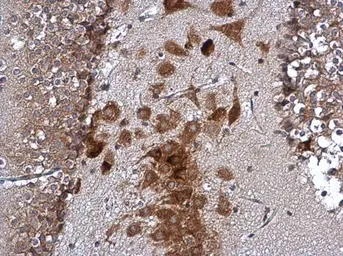
ATXN2 antibody
Cat. No. GTX130329
Cat. No. GTX130329
-
HostRabbit
-
ClonalityPolyclonal
-
IsotypeIgG
-
ApplicationsWB IHC-P IP
-
ReactivityHuman, Mouse