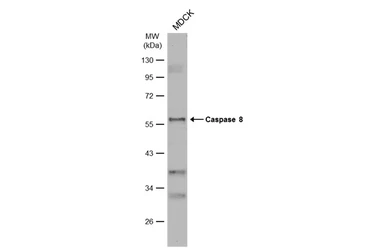
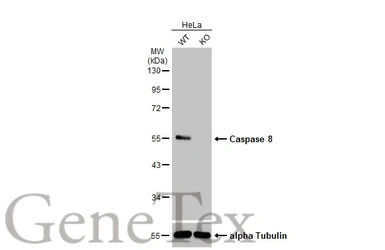
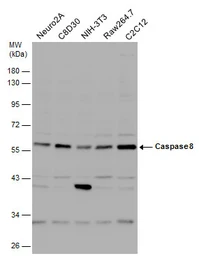
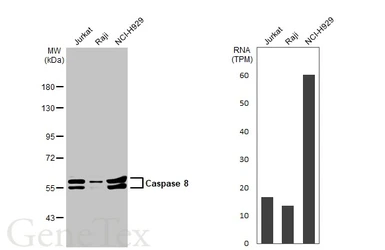
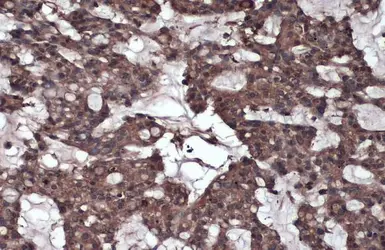
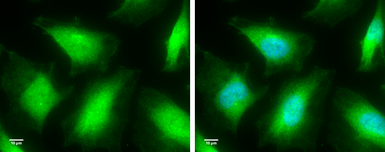
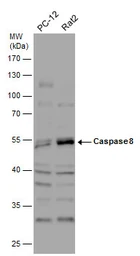
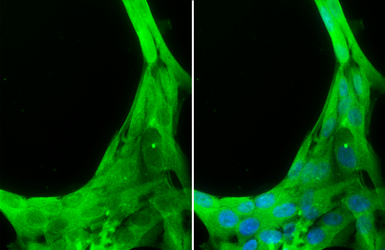
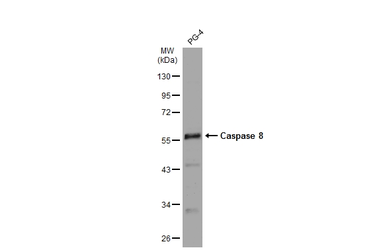
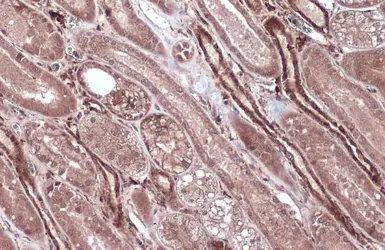
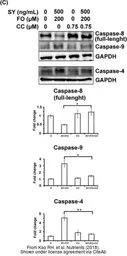
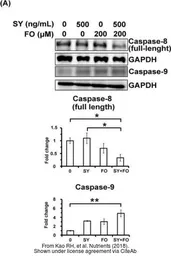
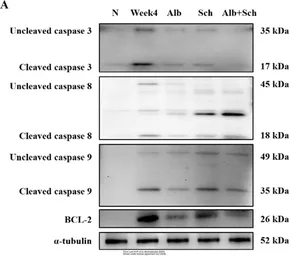
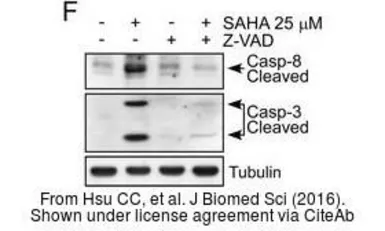
Caspase 8 antibody
Cat. No. GTX110723
Cat. No. GTX110723
-
HostRabbit
-
ClonalityPolyclonal
-
IsotypeIgG
-
ApplicationsWB ICC/IF IHC-P
-
ReactivityHuman, Mouse, Rat, Cat, Dog