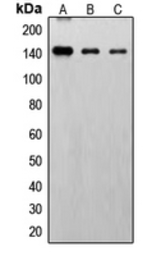
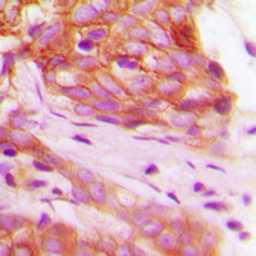
FGFR1 (phospho Tyr766) antibody
Cat. No. GTX32182
Cat. No. GTX32182
-
HostRabbit
-
ClonalityPolyclonal
-
IsotypeIgG
-
ApplicationsWB IHC-P
-
ReactivityHuman, Mouse, Rat